Joint fitting XRT and GBM data with XSPEC models¶
Goals¶
3ML is designed to properly joint fit data from different instruments with thier instrument dependent likelihoods. We demostrate this with joint fitting data from GBM and XRT while simultaneously showing hwo to use the XSPEC models form astromodels
Setup¶
You must have you HEASARC initiated so that astromodels can find the XSPEC libraries.
[2]:
%%capture
import matplotlib.pyplot as plt
from pathlib import Path
from threeML import *
from threeML.io.package_data import get_path_of_data_file
# we will need XPSEC models for extinction
from astromodels.xspec import *
from astromodels.xspec.xspec_settings import *
Load XRT data¶
Make a likelihood for the XRT including all the appropriate files
[4]:
trigger = "GRB110731A"
dec = -28.546
ra = 280.52
p = Path("datasets/xrt")
xrt = OGIPLike(
"XRT",
observation=get_path_of_data_file(p / "xrt_src.pha"),
background=get_path_of_data_file(p / "xrt_bkg.pha"),
response=get_path_of_data_file(p / "xrt.rmf"),
arf_file=get_path_of_data_file(p / "xrt.arf"),
)
fig = xrt.view_count_spectrum()
ax = fig.get_axes()[0]
_ = ax.set_xlim(1e-1)
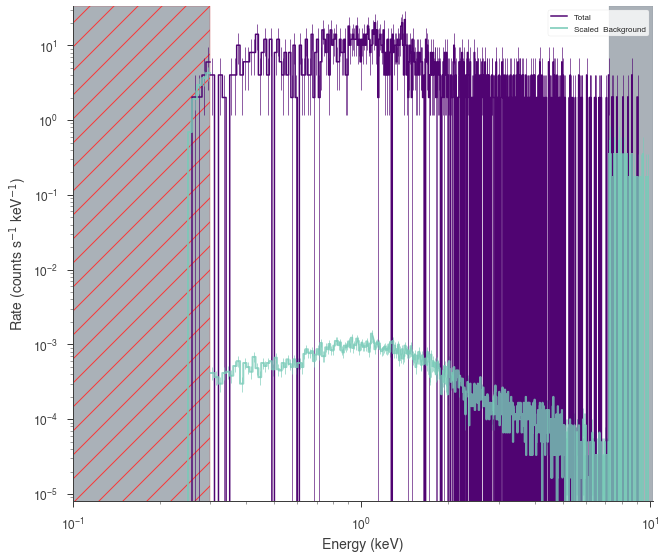
[5]:
fit = xrt.view_count_spectrum(scale_background=False)
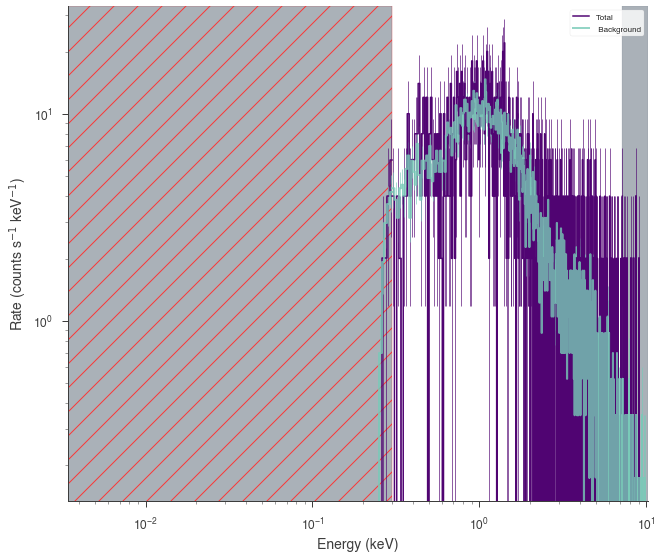
Load GBM data¶
Load all the GBM data you need and make appropriate background, source time, and energy selections. Make sure to check the light curves!
[6]:
trigger_number = "bn110731465"
gbm_data = download_GBM_trigger_data(trigger_number, detectors=["n3"])
[7]:
# Select the time interval
src_selection = "100.169342-150.169342"
bkg_selection = ["-25.0--10.0", "300-400"]
ts = TimeSeriesBuilder.from_gbm_cspec_or_ctime(
name="gbm_n3",
cspec_or_ctime_file=gbm_data["n3"]["cspec"],
rsp_file=gbm_data["n3"]["rsp"],
)
ts.set_background_interval(*bkg_selection)
ts.save_background("n3_bkg.h5", overwrite=True)
fig = ts.view_lightcurve(-50, 450)
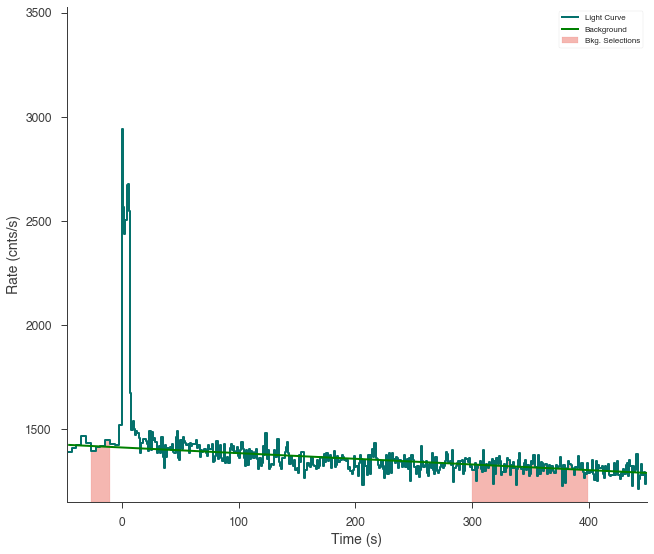
[8]:
ts = TimeSeriesBuilder.from_gbm_tte(
"gbm_n3",
tte_file=gbm_data["n3"]["tte"],
rsp_file=gbm_data["n3"]["rsp"],
restore_background="n3_bkg.h5",
)
ts.set_active_time_interval(src_selection)
fig = ts.view_lightcurve(90, 160)
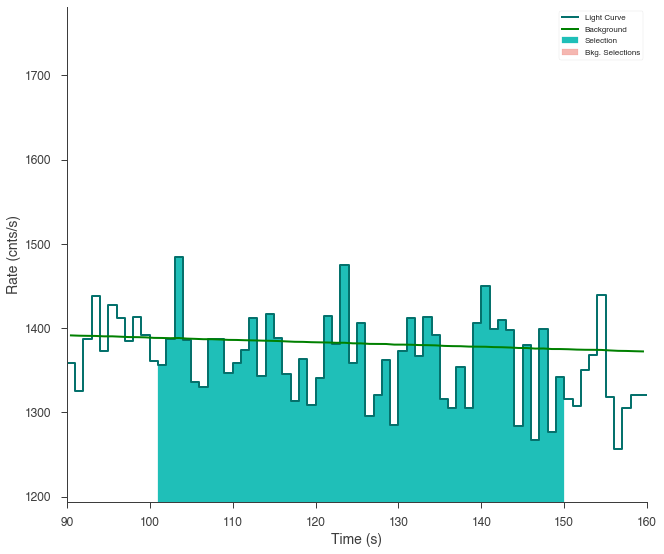
[9]:
nai3 = ts.to_spectrumlike()
Make energy selections and check them out
[10]:
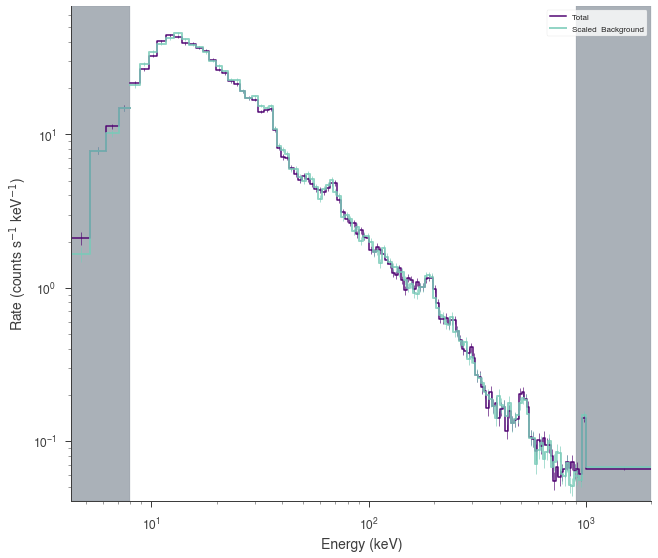
nai3.set_active_measurements("8-900")
fig = nai3.view_count_spectrum()
Setup the model¶
astromodels allows you to use XSPEC models if you have XSPEC installed. Set all the normal parameters you would in XSPEC and build a model the normal 3ML/astromodels way! Here we will use the phabs model from XSPEC and mix it with powerlaw model in astromodels.
With XSPEC¶
[11]:
xspec_abund("angr")
spectral_model = XS_phabs() * XS_zphabs() * Powerlaw()
spectral_model.nh_1 = 0.101
spectral_model.nh_1.bounds = (None, None)
spectral_model.nh_1.fix = True
spectral_model.nh_2 = 0.1114424
spectral_model.nh_2.fix = True
spectral_model.nh_2.bounds = (None, None)
spectral_model.redshift_2 = 0.618
spectral_model.redshift_2.fix = True
Solar Abundance Vector set to angr: Anders E. & Grevesse N. Geochimica et Cosmochimica Acta 53, 197 (1989)
With astromodels PHABS¶
We can build the exact same models in native astromodels thanks to Dominique Eckert. Here, there is no extra function for redshifting the absorption model, just pass a redshift.
[12]:
phabs_local = PhAbs(NH=0.101)
phabs_local.NH.fix = True
phabs_local.redshift.fix = True
phabs_src = PhAbs(NH=0.1114424, redshift=0.618)
phabs_src.NH.fix = True
phabs_src.redshift.fix = True
pl = Powerlaw()
spectral_model_native = phabs_local * phabs_src * pl
Setup the joint likelihood¶
Create a point source object and model.
Load the data into a data list and create the joint likelihood
With XSPEC models¶
First we will fit with the XSPEC model
[13]:
ptsrc = PointSource(trigger, ra, dec, spectral_shape=spectral_model)
model = Model(ptsrc)
[14]:
data = DataList(xrt, nai3)
jl = JointLikelihood(model, data, verbose=False)
model.display()
| N | |
|---|---|
| Point sources | 1 |
| Extended sources | 0 |
| Particle sources | 0 |
Free parameters (2):
| value | min_value | max_value | unit | |
|---|---|---|---|---|
| GRB110731A.spectrum.main.composite.K_3 | 1.0 | 0.0 | 1000.0 | keV-1 s-1 cm-2 |
| GRB110731A.spectrum.main.composite.index_3 | -2.0 | -10.0 | 10.0 |
Fixed parameters (8):
(abridged. Use complete=True to see all fixed parameters)
Linked parameters (0):
(none)
Independent variables:
(none)
[15]:
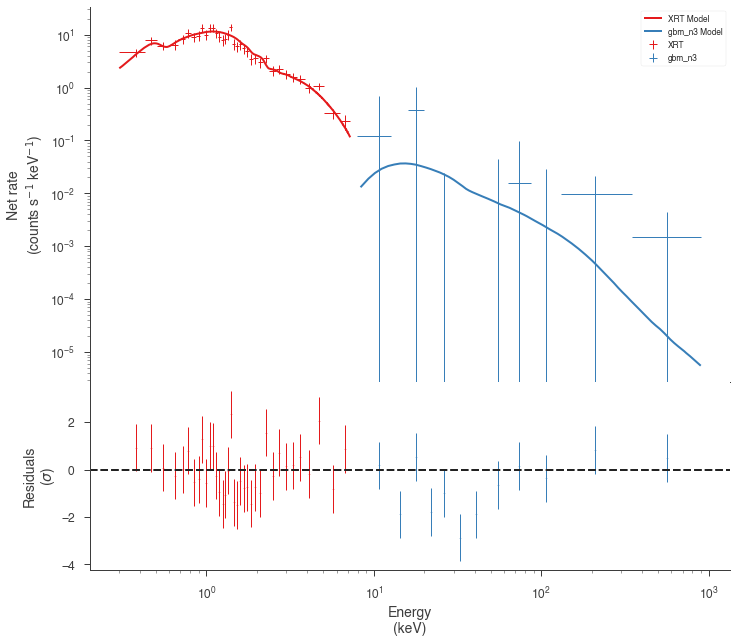
res = jl.fit()
fig = display_spectrum_model_counts(jl, min_rate=[0.5, 0.1])
Best fit values:
| result | unit | |
|---|---|---|
| parameter | ||
| GRB110731A.spectrum.main.composite.K_3 | (1.83 +/- 0.07) x 10^-1 | 1 / (cm2 keV s) |
| GRB110731A.spectrum.main.composite.index_3 | -1.93 +/- 0.05 |
Correlation matrix:
| 1.00 | -0.57 |
| -0.57 | 1.00 |
Values of -log(likelihood) at the minimum:
| -log(likelihood) | |
|---|---|
| XRT | 2064.144639 |
| gbm_n3 | 984.434512 |
| total | 3048.579151 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 6101.173191 |
| BIC | 6110.549900 |
[16]:
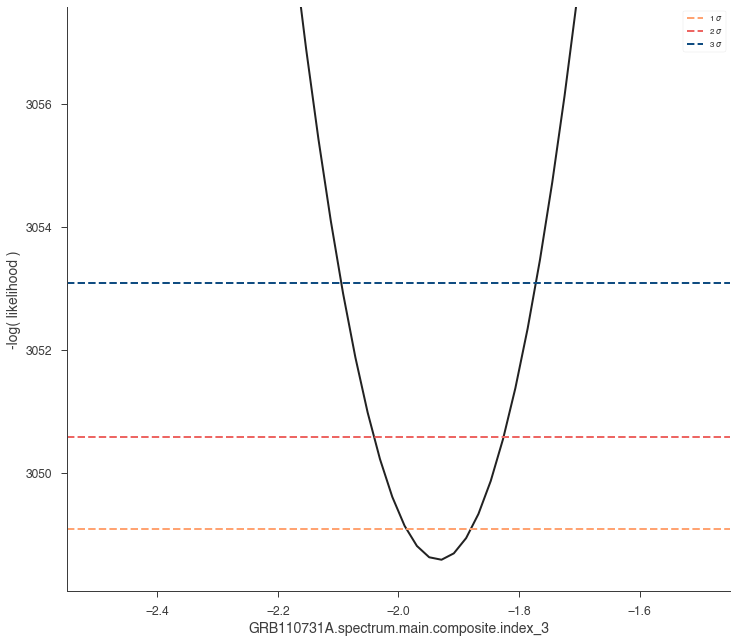
res = jl.get_contours(spectral_model.index_3, -2.5, -1.5, 50)
[17]:
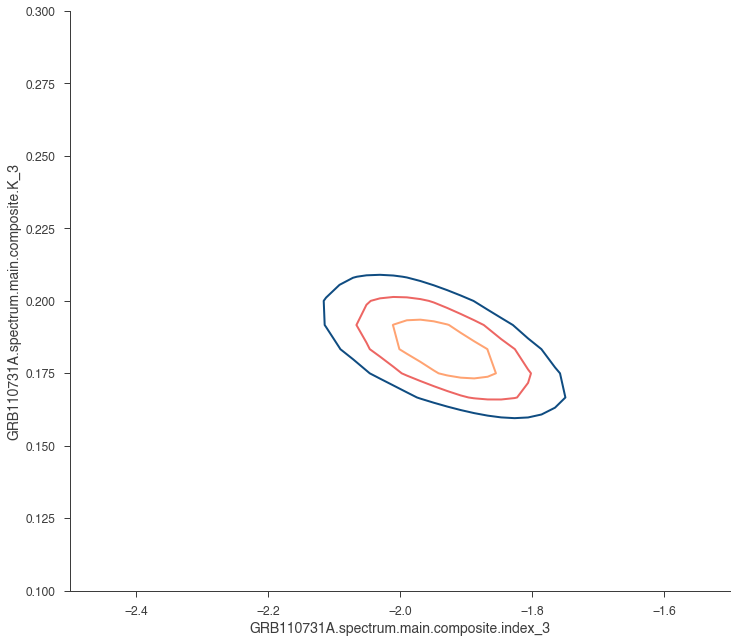
_ = jl.get_contours(
spectral_model.K_3, 0.1, 0.3, 25, spectral_model.index_3, -2.5, -1.5, 50
)
[18]:
fig = plot_spectra(jl.results, show_legend=False, emin=0.01 * u.keV)
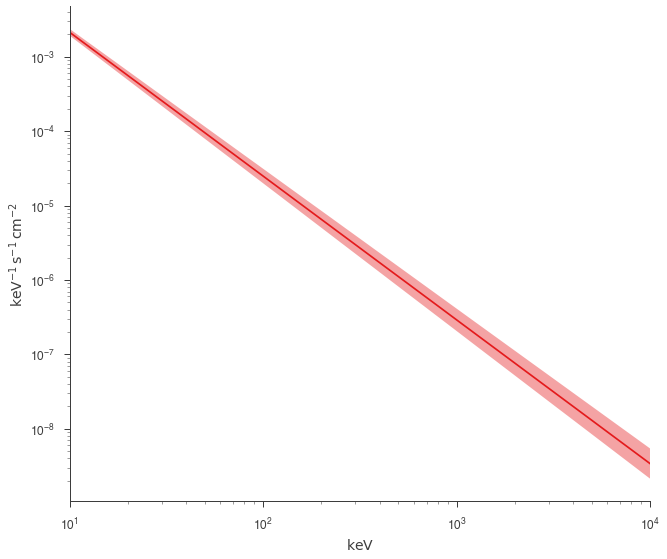
Fit with astromodels PhAbs¶
Now lets repeat the fit in pure astromodels.
[19]:
ptsrc_native = PointSource(trigger, ra, dec, spectral_shape=spectral_model_native)
model_native = Model(ptsrc_native)
[20]:
data = DataList(xrt, nai3)
jl_native = JointLikelihood(model_native, data, verbose=False)
model.display()
| N | |
|---|---|
| Point sources | 1 |
| Extended sources | 0 |
| Particle sources | 0 |
Free parameters (2):
| value | min_value | max_value | unit | |
|---|---|---|---|---|
| GRB110731A.spectrum.main.composite.K_3 | 0.183133 | 0.0 | 1000.0 | keV-1 s-1 cm-2 |
| GRB110731A.spectrum.main.composite.index_3 | -1.933121 | -10.0 | 10.0 |
Fixed parameters (8):
(abridged. Use complete=True to see all fixed parameters)
Linked parameters (0):
(none)
Independent variables:
(none)
[21]:
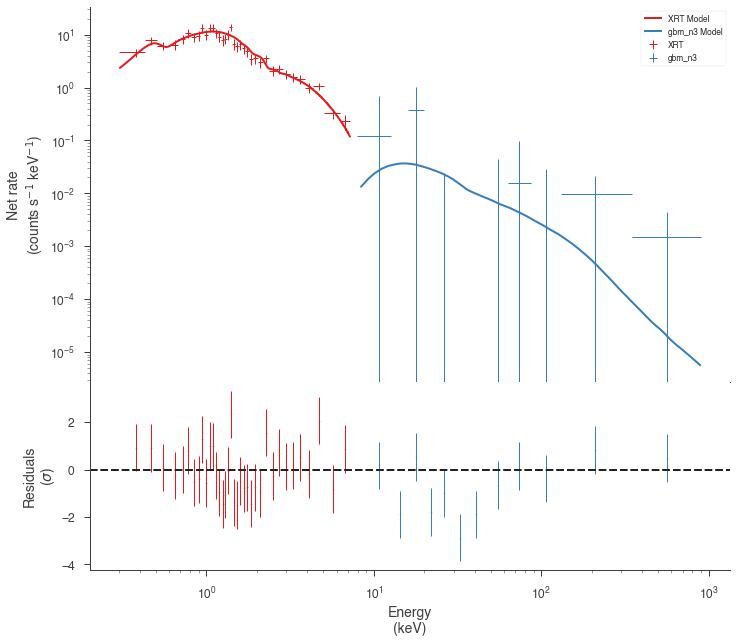
res = jl_native.fit()
fig = display_spectrum_model_counts(jl_native, min_rate=[0.5, 0.1])
Best fit values:
| result | unit | |
|---|---|---|
| parameter | ||
| GRB110731A.spectrum.main.composite.K_3 | (1.83 +/- 0.07) x 10^-1 | 1 / (cm2 keV s) |
| GRB110731A.spectrum.main.composite.index_3 | -1.93 +/- 0.05 |
Correlation matrix:
| 1.00 | -0.57 |
| -0.57 | 1.00 |
Values of -log(likelihood) at the minimum:
| -log(likelihood) | |
|---|---|
| XRT | 2064.134895 |
| gbm_n3 | 984.434410 |
| total | 3048.569305 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | 6101.153498 |
| BIC | 6110.530208 |
[22]:
fig = plot_spectra(jl.results, jl_native.results, show_legend=False, emin=0.01 * u.keV)
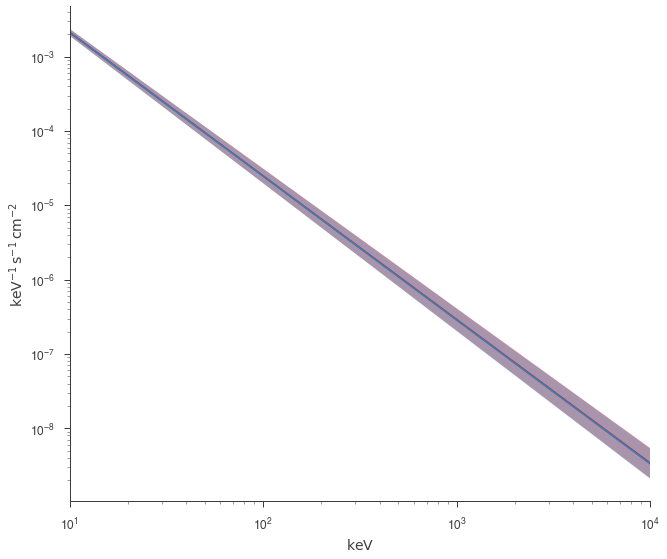
Both approaches give the same answer as they should.
[ ]: